 You can see how many features were counted on the basis of information you provided in SAF table. fc_SE <- featureCounts("alignResults.BAM",annot.ext=ann) BAM file that we created by aligning to Senecavirus A genome have accession number liked to each reads. Here you have to use Genebank accession number of virus genome as Chr. So, lets use follwing code to create one SAF file for this analysis. Rsubread package allows you to create such files in tabular format which they call it SAF format. For viruses, in most of the cases you don’t find well-annotated GTF or GFF files. We need a annotated file in GTF or GFF format to count the features or genes aligned. I saw that all the reads that were in meta.fastq belonging to Senecavirus A were aligned while, all the Zika virus reads were ignored.The output will be in. align(index="seneca",readfile1=”meta.fastq”,output_file="alignResults.BAM") Now, I can align reads in meta.fastq to the index file which I created above. In my case it would be buildindex(basename= “seneca”, reference= “sva.fasta”) #syntaxīuildindex(basename="reference_index",reference=ref) Use the same Senecavirus A whole genome file which you used to extract reads in the above example. First, we need to build index of our reference files.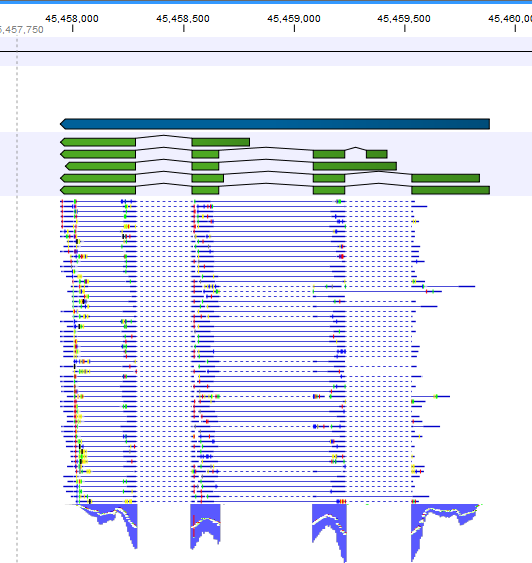 
#meta.fasta is my input file, meta.fastq is the output file and we are assigning quality score of 34 to all the basepairs. reformat.sh in= meta.fasta out=meta.fastq qfake=35 You can find details about bbmap and reformat.sh script elsewhere. I used reformat.sh script which is a part of bbmap. There are many tools available to convert fasta file to fastq format. This fasta file needs to be changed into fastq format. TGCCGTACCGAGTCACGAGTACCTGCAGGCAAGATGGAGGGCCTTGTTCGACTGACCTGGATAGCCCAACGCGCTTCGGTGCTGCCGGCGATTCTGGGAGAACTCAGTCGGAĪGTTGTTGATCTGTGTGAATCAGACTGCGACAGTTCGAGTTTGAAGCGAAAGCTAGCAACAGTATCAACAĪAAGCTAGCAACAGTATCAACAGGTTTTATTTTGGATTTGGAAACGAGAGTTTCTGGTCATGAAAAACCCA GTGTGGTCTGCGAGTTCTAGCCTACTCGTTTCTCCCCTACTCACTCATTCACACACAAAAAĬTACAAGATTTGGCCCTCGCACGGGATGTGCGATAACCGCAAGATTGACTCAAGCGCGGAAAGCGCTGTAACC GTTTCTCCCCTACTCACTCATTCACACACAAAAACTGTGTTGTAACTACAAGATTTGGCCCTCGCACGGGĪTGTGCGATAACCGCAAGATTGACTCAAGCGCGGAAAGCGCTGTAACCACATGCTGTTAGTCCCTTTATGĬGGGGGGTAAACCGGCTGTGTTTGCTAGAGGCACAGAGGAGCAACATCCAACCTGCTTTTGTĬGGCTCCAATTCCTGCGTCGCCAAAGGTGTTAGCGCACCCAA So, Zika virus reads should not be counted by Rsubread while aligning.ĪAGGAAGGACTGGGCATGAGGGCCCAGTCCTTCCTTTCCCCTTCCGGGGGGTAAACCGGCTGTGTTTGCTĪGAGGCACAGAGGAGCAACATCCAACCTGCTTTTGTGGGGAACGGTGCGGCTCCAATTCCTGCGTCGCCAĪAGGTGTTAGCGCACCCAAACGGCGCATCTACCAATGCTATTGGTGTGGTCTGCGAGTTCTAGCCTACTC I will be aligning my reads to Senecavirus A genome. And I also pulled some sequences from the Zika virus which are names as Zika1 and Zika2. I created a fasta file with a few contigs each containing about 70-100 basepairs, and named each contig as read 1, read 2 and so on. fasta file by pulling some of the sequences from the Senecavirus A genome. 
source("")įor this simulation I created a small. Another important aspect of learning RNA-Seq analysis is understanding the algorithms behind the analysis.To this end, I decided to run a small simulation to understand how RNA-Seq analysis algorithms work.It is amazing how a single R package can do things like read aligning, read mapping and read counts in few lines of codes. Softwares with graphical user interface like CLC Workbench, have made RNA-Seq data analysis quite easier.However, they are expensive and in most of the cases you might not be able to tweak your analysis in the exact way you want. RNA-Seq data analysis can be complicated. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
